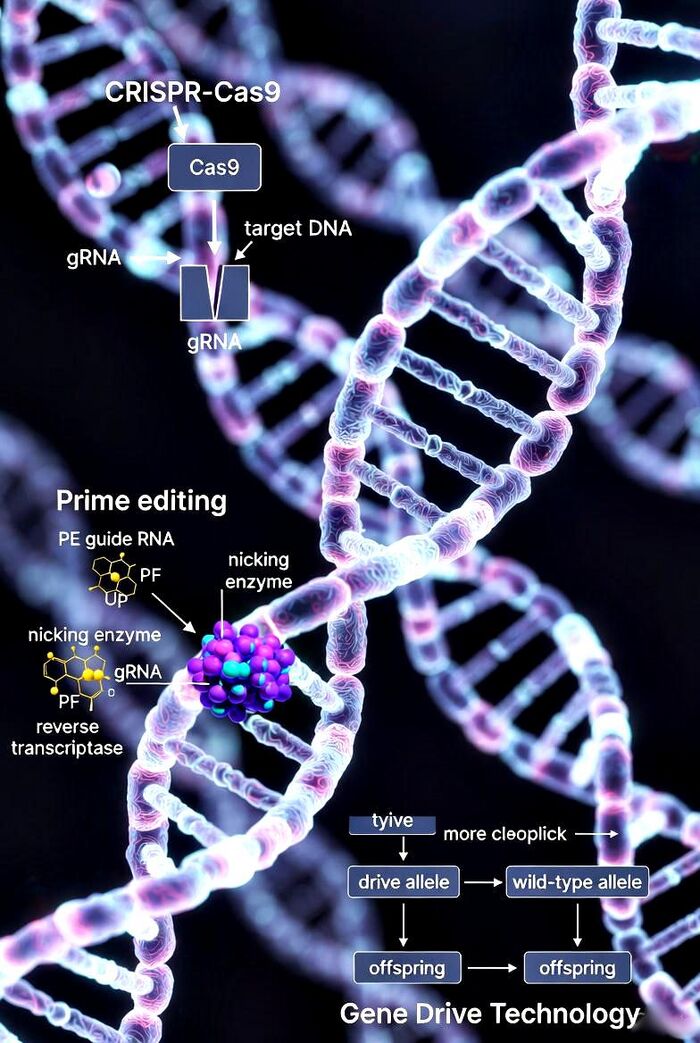
CRISPR-Cas9
CRISPR-Cas9 (Clustered Regularly Interspaced Short Palindromic Repeats, CRISPR-associated protein 9) is a groundbreaking gene-editing technology that enables precise modifications to an organism's DNA. Derived from a natural bacterial defense mechanism against viruses, CRISPR-Cas9 functions as molecular scissors, allowing scientists to cut, edit, and insert genetic material with unprecedented accuracy and efficiency. This tool has revolutionized the field of Genome Editing, transforming it from a labor-intensive process into a rapid, accessible method for altering genomes across a wide range of organisms, from bacteria to humans.
At its core, CRISPR-Cas9 exemplifies the intersection of evolutionary biology and synthetic engineering. In bacteria, the CRISPR system stores snippets of viral DNA as "spacers" between repetitive sequences, enabling the Cas9 enzyme—guided by a complementary RNA molecule—to recognize and cleave invading viral genomes. Researchers, led by pioneers such as Jennifer Doudna and Emmanuelle Charpentier, adapted this system in 2012 to target specific DNA sequences in eukaryotic cells, earning them the 2020 Nobel Prize in Chemistry. This adaptation has made genome editing not only feasible but scalable, democratizing access to genetic manipulation for applications in medicine, agriculture, and basic research.
The technology's simplicity—requiring only a guide RNA (gRNA) to direct Cas9 to the target site—contrasts sharply with earlier methods like zinc-finger nucleases or TALENs, which demanded complex protein engineering. In genome editing, CRISPR-Cas9 introduces double-strand breaks at precise locations, triggering cellular repair mechanisms such as non-homologous end joining (NHEJ) for gene disruption or homology-directed repair (HDR) for precise insertions or corrections. This versatility underpins its dual potential: immense promise for curing genetic diseases and profound risks of misuse, from unintended off-target mutations to ethical dilemmas in human germline editing.
By December 2025, CRISPR-Cas9 has matured into a multifaceted platform, with over 100 clinical trials underway worldwide. Recent milestones include the approval of personalized in vivo therapies for rare diseases, such as a bespoke treatment for carbamoyl-phosphate synthetase 1 (CPS1) deficiency in an infant, developed in just six months. These advances underscore CRISPR's role in precision medicine, while ongoing refinements in delivery systems, like lipid nanoparticles, address key barriers to broader adoption.
History
The discovery of CRISPR sequences dates back to 1987, when Japanese researchers Osamu Shimohara and Yoshizumi Ishino identified unusual repeating DNA patterns in the Escherichia coli genome while studying alkaline phosphatase. These patterns, initially dismissed as regulatory elements, were later recognized in 2002 by Francisco Mojica as remnants of phage infections, forming an adaptive immune system in prokaryotes. Concurrent work by Gilles Vergnaud and others mapped similar arrays in Mycobacterium tuberculosis, solidifying the CRISPR concept.
Key breakthroughs occurred in the early 2000s. In 2005, three independent teams—led by Mojica, Alexander Bolotin, and Ruud Jansen—simultaneously proposed that CRISPR spacers matched viral sequences, suggesting a memory function. By 2007, Phillip Horvath and Rodolphe Barrangou at Danisco demonstrated CRISPR's role in conferring bacterial resistance to bacteriophages, coining the term "CRISPR" in a seminal paper. The identification of Cas genes, including Cas9 from Streptococcus pyogenes, followed in 2008, revealing the machinery for spacer acquisition, processing, and interference.
The leap to genome editing came in 2012, when Doudna and Charpentier engineered a simplified system using CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA) fused into a single guide RNA, paired with Cas9 for in vitro DNA cleavage. This "CRISPR 2.0" was rapidly adapted for cellular use by Feng Zhang and George Church's groups in 2013, enabling targeted edits in mammalian cells. Subsequent refinements, such as Cas9 variants for nickase activity or base editing, have expanded its precision.
Entering the 2020s, CRISPR's commercialization accelerated with the 2023 FDA approval of Casgevy for sickle cell disease and beta-thalassemia, marking the first CRISPR-based therapy. By 2025, advancements include CRISPR-Cas12a for multiplexed immunological modeling and enhanced specificity tools like eSpCas9 and HypaCas9, reducing off-target effects. These developments have propelled CRISPR into routine use in drug discovery and personalized medicine.
The following table outlines key categories of historical events in CRISPR-Cas9 development, highlighting its evolution within the broader context of genome editing.
The following table summarizes major milestones in CRISPR-Cas9 history.
| Category | Event | Historical Context | Initial Promotion as Science | Emerging Evidence and Sources | Current Status and Impacts |
|---|---|---|---|---|---|
| Discovery | Identification of CRISPR repeats | Late 1980s: Routine sequencing of bacterial genes reveals palindromic repeats | Viewed as novel regulatory motifs | 1987 paper on E. coli genome; later spacer analysis in 2002 | Foundation for bacterial immunity studies; precursor to editing tech |
| Functional Insight | Spacer-protospacer matching | Early 2000s: Phage-bacteria coevolution studies | Proposed as viral defense memory | 2005 papers by Mojica, Bolotin, Jansen | Established CRISPR as adaptive immunity; inspired biotech applications |
| Enzymatic Role | Cas9 characterization | Mid-2000s: Yogurt industry phage resistance research | Demonstrated as targeted nuclease | 2007 Barrangou-Horvath paper in Science | Commercial use in food safety; bridge to programmable editing |
| Engineering Breakthrough | In vitro CRISPR-Cas9 system | 2010s: Rise of synthetic biology | Hyped as "genetic scissors" for universal editing | 2012 Doudna-Charpentier Science paper | Nobel recognition in 2020; explosion in genome editing tools |
| Eukaryotic Application | Mammalian cell editing | 2013: CRISPR race among labs | Promoted for disease modeling and therapy | Zhang and Church Nature/Science papers | Thousands of publications; clinical trials initiated |
| Therapeutic Milestone | Casgevy approval | 2020s: Precision medicine surge | Framed as curative for blood disorders | 2023 FDA approval; 2025 UK NICE endorsement | First commercial CRISPR therapy; expands to 100+ trials |
| Personalization Advance | Bespoke in vivo therapy | 2025: Rare disease focus | Touted for on-demand editing | CHOP/Penn Medicine infant treatment for CPS1 deficiency | Sets regulatory precedent; accelerates rare disease pipelines |
Mechanism
CRISPR-Cas9 operates through a multi-step process that integrates RNA-guided DNA recognition with endonuclease activity, making it a cornerstone of modern genome editing. The system comprises two main components: the Cas9 protein, a RNA-guided DNA endonuclease, and a guide RNA (gRNA) that directs it to the target.
Upon expression in a target cell, the gRNA—typically a chimeric molecule combining crRNA and tracrRNA—binds to complementary DNA sequences adjacent to a protospacer adjacent motif (PAM), usually 5'-NGG-3' for S. pyogenes Cas9. This binding induces a conformational change in Cas9, enabling it to scan and hybridize with the target DNA, displacing the non-target strand to form an RNA-DNA hybrid. Cas9 then cleaves both strands ~3-4 base pairs upstream of the PAM, generating a blunt-ended double-strand break (DSB).
In the context of genome editing, this DSB activates endogenous repair pathways. NHEJ often introduces small insertions or deletions (indels) at the break site, leading to frameshift mutations and gene knockout. For precise edits, a donor template with homology arms can be provided to exploit HDR, allowing gene correction, insertion, or replacement. Off-target effects, where Cas9 cuts unintended sites with partial gRNA complementarity, remain a challenge, mitigated by high-fidelity Cas9 variants or prime editing strategies.
This mechanism's programmability—simply altering the gRNA sequence—distinguishes CRISPR from restriction enzymes, enabling multiplexed editing of multiple loci simultaneously and paving the way for large-scale genomic rewrites. Recent 2025 innovations, such as cell-permeable protein controls for Cas9 deactivation, further enhance temporal precision, minimizing prolonged exposure and off-target risks.
Genome Editing
Genome editing refers to the deliberate alteration of an organism's genetic material, encompassing techniques to add, delete, or modify DNA sequences. CRISPR-Cas9 has emerged as the gold standard in this field, supplanting older tools due to its ease, cost-effectiveness, and specificity. Prior methods, such as meganucleases (1980s), zinc-finger nucleases (ZFNs, 1990s), and transcription activator-like effector nucleases (TALENs, 2010s), required bespoke protein design for each target, limiting scalability.
CRISPR democratizes genome editing by allowing off-the-shelf gRNA synthesis, reducing design time from months to days. Applications span from creating knockout libraries for functional genomics to engineering synthetic chromosomes. In research, it facilitates loss-of-function screens to identify gene roles; in therapy, it targets monogenic disorders like sickle cell anemia via ex vivo editing of hematopoietic stem cells.
However, genome editing with CRISPR raises technical hurdles, including delivery challenges (e.g., viral vectors for in vivo use) and epigenetic considerations, as edits may not fully recapitulate natural variation. Nonetheless, its integration with technologies like single-cell sequencing and CRISPR interference (CRISPRi) for transcriptional control expands genome editing into epigenome modulation and beyond. By 2025, mRNA-driven CRISPR deliveries have addressed immunogenicity issues, enabling safer in vivo applications for autoimmune and oncological diseases.
Uses
The applications of CRISPR-Cas9 are vast, spanning therapeutic, agricultural, and industrial domains, all rooted in its prowess for precise genome editing. In medicine, it holds transformative potential for treating genetic diseases. For instance, editing the CCR5 gene has rendered cells resistant to HIV in proof-of-concept studies, while clinical trials using CRISPR to correct BCL11A mutations in beta-thalassemia and sickle cell disease have yielded transfusion-independent patients, as seen in Vertex Pharmaceuticals' Casgevy therapy approved in 2023 and expanded in 2025.
Beyond monogenic disorders, CRISPR targets complex conditions like cancer by engineering CAR-T cells with enhanced tumor specificity or disrupting PD-1 checkpoints. In neurology, it addresses Huntington's disease by silencing mutant HTT alleles, and in ophthalmology, it restores vision in Leber congenital amaurosis via subretinal injections. Emerging uses include epigenetic editing to reactivate silenced genes in Alzheimer's models. Cardiovascular applications, such as Verve Therapeutics' (acquired by Eli Lilly in 2025) in vivo editing for high cholesterol, demonstrate CRISPR's reach into polygenic traits.
Agriculturally, CRISPR enhances crop resilience and yield. Gene-edited mushrooms resistant to browning (2016 USDA approval) bypassed GMO regulations, while drought-tolerant rice and virus-resistant cassava address food security. Livestock improvements, such as hornless cattle via PCK1 knockout, reduce animal welfare issues without transgenes. Gene drive technology, leveraging CRISPR to bias inheritance and spread traits through populations, offers revolutionary pest control; for example, engineered Anopheles gambiae mosquitoes suppress malaria transmission by driving sterility genes.
In basic science, CRISPR enables high-throughput screens, evolutionary studies via directed evolution, and microbiome engineering for gut health. Industrially, it optimizes microbial strains for biofuel production or pharmaceutical synthesis, such as insulin from edited yeast. Diagnostic tools like SHERLOCK use Cas13 for rapid pathogen detection, amplified in 2025 with SNV-specific CRISPRdx for cancer monitoring.
The following table categorizes key uses of CRISPR-Cas9, emphasizing its role in advancing genome editing paradigms.
The following table provides an overview of CRISPR-Cas9 applications by category.
| Category | Event | Historical Context | Initial Promotion as Science | Emerging Evidence and Sources | Current Status and Impacts |
|---|---|---|---|---|---|
| Therapeutic | Sickle cell editing trials | 2010s: Rise of precision medicine | Touted as cure for inherited blood disorders | 2017 first human trial; 2023 FDA approval of Casgevy | Over 50 trials; potential for 7,000+ rare diseases |
| Agricultural | Non-browning mushrooms | 2010s: GMO debates intensify | Promoted as "natural" trait editing | 2015 Penn State development; 2016 deregulation | Expanded to 20+ crops; boosts nutrition and yield |
| Research | Genome-wide knockout screens | 2014: Big data genomics era | Hyped for functional annotation of genomes | Shalem et al. Science 2014 | Millions of variants mapped; accelerates drug discovery |
| Industrial | Biofuel strain optimization | 2010s: Synthetic biology boom | Marketed for sustainable manufacturing | 2016 CRISPR-edited algae for lipids | Commercial pilots; reduces fossil fuel reliance |
| Microbiome | Gut bacteria engineering | Late 2010s: Microbiota research surge | Positioned for personalized probiotics | 2018 studies on Bacteroides editing | Preclinical; promises for IBS and obesity treatment |
| Ecological | Gene drive in mosquitoes | 2010s: Vector-borne disease crises | Hyped for malaria eradication | 2015 Anopheles lab success; 2025 field trials | Population suppression; 90%+ trait spread in models |
| Diagnostic | SHERLOCK for pathogens | 2017: Point-of-care testing rise | Framed as portable lab replacement | 2025 SNV-specific enhancements | Attomolar sensitivity; deployed in outbreaks |
Abuses and Ethical Concerns
While CRISPR-Cas9's uses herald a new era in genome editing, its accessibility amplifies risks of abuse, from biosecurity threats to socioeconomic disparities. Ethically, the technology's power to alter heritable genomes evokes eugenics fears, as evidenced by the 2018 scandal involving He Jiankui, who used CRISPR to create HIV-resistant babies by editing CCR5 in embryos—condemned globally for safety lapses and consent violations, resulting in his imprisonment.
Misuses include dual-use research of concern (DURC), where editing enhances pathogen virulence, such as hypothetical gain-of-function in influenza. Non-state actors could engineer bioweapons, prompting calls for international moratoriums. Off-target mutations pose oncogenic risks in therapies, with studies reporting up to 5% unintended edits, necessitating improved specificity. Gene drives exacerbate these concerns: while aimed at eradicating pests, unintended spread to non-target species could cause ecological collapses, as seen in modeling of collateral damage in mosquito complexes.
Equity issues loom large: patent battles (e.g., Broad Institute vs. UC Berkeley) restrict access in developing nations, exacerbating the "genomic divide." Designer babies, selecting traits like intelligence via polygenic editing, challenge human dignity, while environmental releases of gene drives risk biodiversity loss and resistance evolution. Sociological risks include public backlash and governance gaps, with 2025 reviews highlighting malevolent uses like terrorist-driven drives for harmful traits.
Regulatory responses vary: the U.S. classifies most CRISPR edits as non-GMO if no foreign DNA is added, while the EU imposes stringent oversight. Global frameworks, like the 2018 WHO gene editing report updated in 2025, advocate for inclusive governance to balance innovation with precaution, including ecological risk assessments for drives.
The following table delineates categories of potential abuses, contextualizing them within genome editing's ethical landscape.
The following table explores abuses and concerns associated with CRISPR-Cas9.
| Category | Event | Historical Context | Initial Promotion as Science | Emerging Evidence and Sources | Current Status and Impacts |
|---|---|---|---|---|---|
| Germline Editing | He Jiankui's CRISPR babies | 2010s: IVF advancements meet editing tech | Framed as humanitarian HIV prevention | 2018 revelation; embryo CCR5 edits | International summits; de facto bans in 70+ countries |
| Dual-Use Risks | Pathogen enhancement experiments | Post-9/11 biosecurity focus | Defended as gain-of-insights for vaccines | 2011 H5N1 airborne studies (pre-CRISPR analog) | U.S. DURC oversight; export controls on tools |
| Off-Target Effects | Mosaic mutations in trials | 2010s: Early human applications | Minimized as rare artifacts | 2018 Nature Medicine analysis of cancer risks | High-fidelity variants; required in trial protocols |
| Equity Disparities | Patent disputes limiting access | 2010s: Biotech commercialization | Promoted as global good | 2014-ongoing Broad vs. Berkeley litigation | Open-source initiatives; calls for tiered pricing |
| Ecological Misuse | Gene drive releases | 2010s: Vector-borne disease crises | Hyped for malaria eradication | 2015 Anopheles drive success in labs | Field trials paused; biodiversity impact assessments |
| Biosecurity Threats | Engineered bioweapons | 2020s: DIY bio labs proliferation | Warned as terrorist tools | 2025 reviews on malevolent drives | Enhanced surveillance; international treaties proposed |
Variants and Future Directions
Evolving from the original Cas9, CRISPR variants like Cas12 and Cas13 expand editing to RNA targets or collateral cleavage for diagnostics (e.g., SHERLOCK). Base editors enable single-nucleotide changes without DSBs, reducing indels, while prime editing allows versatile insertions up to 44 bp. Future trajectories include in vivo delivery via nanoparticles and integration with AI for gRNA design, further embedding CRISPR in genome editing's toolkit.
=== Prime Editing ===
Prime editing, developed by David Liu's lab in 2019, represents a pinnacle of CRISPR evolution, enabling "search-and-replace" modifications without double-strand breaks. The system fuses a Cas9 nickase (nCas9, H840A mutation) with a reverse transcriptase (RT) to form the prime editor (PE), guided by a prime editing guide RNA (pegRNA). The pegRNA includes a spacer for target recognition, a reverse transcription template (RTT) encoding the desired edit, and a primer binding site (PBS) for flap hybridization.
Mechanistically, nCas9 nicks the target strand, exposing a 3' flap that hybridizes with the PBS. RT then synthesizes new DNA from the RTT, incorporating edits like point mutations, insertions (up to 44 bp), deletions, or combinations. This flap anneals to the complementary strand, displacing the original, and is resolved by cellular machinery, yielding precise changes with minimal indels or off-targets compared to CRISPR-Cas9.
Applications abound: prime editing corrects cystic fibrosis mutations in preclinical models and enables multiplexed edits for cancer immunotherapy. In 2025, the PERT (prime editing-mediated readthrough of premature termination codons) variant addresses nonsense mutations—causing ~1/3 of genetic diseases—by inserting readthrough sequences, potentially treating diverse conditions with a single agent. Clinical trials for prime-edited therapies in hemoglobinopathies are slated for 2026, with efficiencies reaching 50-70% in human cells via engineered RT domains.
Challenges include pegRNA stability and delivery, but 2025 protein engineering boosts activity 10-fold. Prime editing's scarless precision positions it as a next-generation tool for therapeutic genome editing, complementing CRISPR-Cas9 in complex repairs.
=== Gene Drive Technology ===
Gene drives harness CRISPR-Cas9 to override Mendelian inheritance, ensuring modified alleles spread through populations at rates exceeding 50%. In a homing drive, Cas9 and gRNA target the wild-type allele on the homologous chromosome, cleaving it and copying the drive via homology-directed repair—effectively converting heterozygotes to homozygotes.
Uses focus on population control: suppression drives induce sterility or lethality, as in 2025 lab trials of Aedes aegypti mosquitoes to curb dengue. Replacement drives introduce anti-parasite genes, like those blocking Plasmodium in Anopheles, potentially slashing malaria cases by 90% in models. Conservation applications target invasives, such as rodents on islands, restoring ecosystems.
Risks are profound: drives could hybridize with non-target species, causing unintended extinctions or resistance evolution. 2025 studies highlight collateral damage in mosquito complexes, urging reversible "daisy drives" with kill switches. Deployment requires rigorous modeling and stakeholder consensus, with field trials limited to contained settings.
Challenges persist, such as immune responses to Cas9 and delivery barriers, but optimism abounds for curing polygenic diseases and engineering climate-resilient ecosystems. As genome editing matures, CRISPR-Cas9 remains its linchpin, demanding vigilant stewardship to harness benefits while averting harms. By 2025, SyNTase editing platforms promise scalable corrections, while regulatory pathways for personalized therapies signal a new era.
Categories
To encapsulate the thematic breadth of CRISPR-Cas9, the following categories delineate its multifaceted role in science, society, and ethics, with a focus on genome editing implications.
| Category | Sub-themes | Relation to Genome Editing | Societal Implications | Mitigation Strategies |
|---|---|---|---|---|
| Scientific Innovation | Mechanism refinement, variant development | Enables precise, scarless edits beyond DSBs | Accelerates discoveries in biology and medicine | Open-access repositories for protocols |
| Therapeutic Applications | Disease correction, immunotherapy | Targets causal variants in complex genomes | Reduces healthcare burdens; personalizes treatments | Rigorous clinical trials and pharmacovigilance |
| Agricultural Enhancement | Trait stacking, stress tolerance | Edits polyploid genomes efficiently | Enhances food security amid climate change | Biosafety assessments and farmer education |
| Ethical Dilemmas | Germline modifications, consent issues | Alters heritable traits irreversibly | Risks inequality and unintended societal shifts | International accords and public deliberation |
| Biosecurity Threats | Weaponization potential, accidental releases | Scales editing to novel organisms | Poses global health and environmental risks | Surveillance frameworks and dual-use training |
| Ecological Interventions | Gene drives, biodiversity engineering | Biases population-level edits | Balances pest control with ecosystem stability | Reversible designs and impact modeling |
| Technological Refinements | Prime editing, specificity boosts | Minimizes errors in therapeutic edits | Improves safety for widespread adoption | AI-guided designs and high-fidelity enzymes |